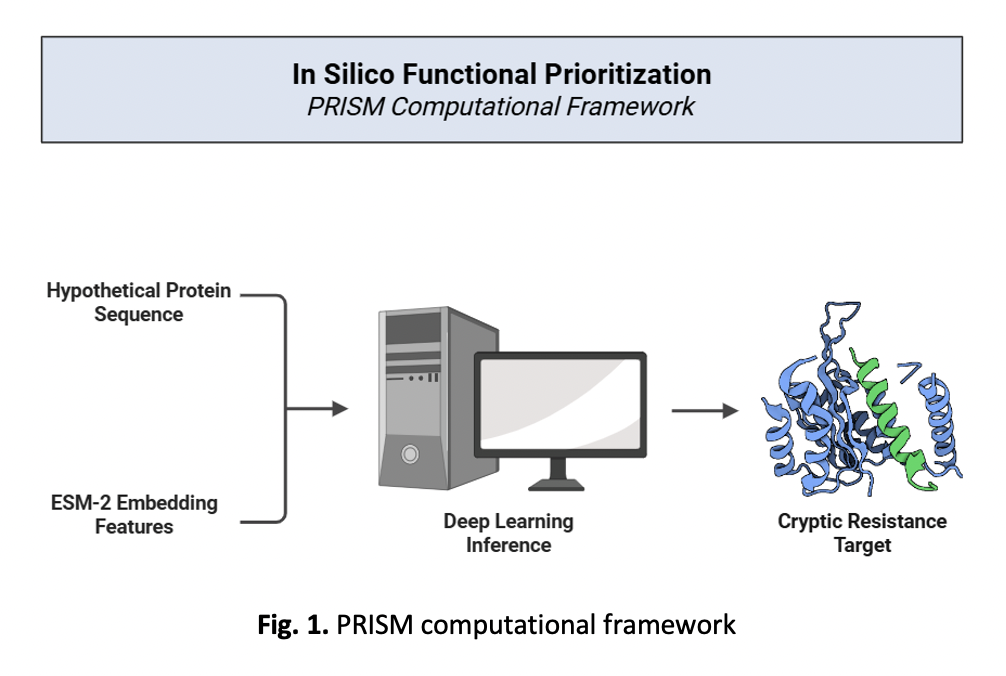
In Silico Functional Prioritization of Hypothetical Proteins Associated with Multidrug Resistance in Acinetobacter baumannii using PRISM Computational Framework
Keywords:
Acinetobacter baumannii, antimicrobial resistance, hypothetical proteins, deep learningAbstract
Acinetobacter baumannii is a critical-priority pathogen whose pervasive multidrug-resistant (MDR) phenotype poses a global clinical crisis. A major challenge in combating this pathogen is the vast number of hypothetical proteins that remain uncharacterized, representing a hidden reservoir of potential resistance mechanisms that evade standard bioinformatics tools. The primary objective of this study is to computationally prioritize these hypothetical proteins to uncover novel therapeutic targets, specifically addressing the limitation of biologically implausible predictions often generated by standard unconstrained deep learning models. To achieve this, we utilized the PRISM (Protein Recognition Insight via Sequence Modelling) framework, which integrates protein language model embeddings with strict biological logic. The analysis revealed twelve cryptic Gcn5-related N-acetyltransferase (GNAT) determinants systematically missed by consensus-based annotation. By enforcing ontological constraints, PRISM reduced the experimental search space by 99.5%, significantly outperforming standard baselines (McNemar’s test, p< 0.001) and drastically reducing biological hallucinations from 15.99% to 0.04%. Topology modelling demonstrated that these enzymes utilize a non-canonical "border security" strategy, confirming their sequestration in the cell membrane or secretion to the periplasm via N-terminal signal peptides. Additionally, our analysis established a novel class of minimalist 65-amino acid resistance proteins and resolved a systemic "metabolic bias" in global repositories by reassigning mislabelled metabolic proteins as secreted defense determinants. Ultimately, these findings redefine the genomic "dark matter" of A. baumannii as a strategically localized reservoir of enzymatic defense, providing a critical, cost-effective corrective layer for antimicrobial surveillance.